December 10, 2019
Logarithmic models
What is a logarithm?
We need very little math for our course: arithmetic, algebra, and logarithms
Just remember that if \(x=p^m\) then \[\log_p(x) = m\]
If we use another base, for example \(q\), then \[\log_q(x) = m\cdot\log_q(p)\]
So if we use different bases, there is only a scale factor
The easiest one is natural logarithm
Other things about logarithms
- They only work with positive numbers. Not with 0
- If \(x=p\cdot q\) then \[\log(x)=\log(p)+\log(q)\]
- If \(x=a^m\) then \[\log(x)=m\log(a)\]
- If \(x=\exp(m)\) then \[\log(x)=m\]
Linear models can be used in three cases
Basic linear model \[y=A+B\cdot x\] Exponential \[y=I\cdot R^x\qquad\log(y)=log(I)+log(R)\cdot x\] Power of \(x\) \[y=C\cdot x^E\qquad\log(y)=log(C)+E\cdot\log(x)\]
Which one to use?
The easiest way to decide is to
- draw several plots, placing
log()in different places, - see which one seems more like a straight line
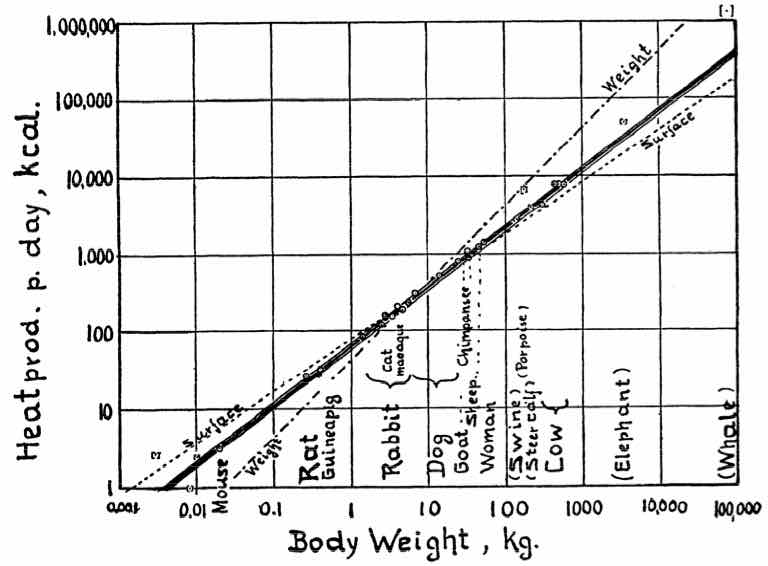
For example, let’s analyze data from Kleiber’s Law
The following data shows a summary. The complete table has 26 animals
Body size v/s metabolic rate
| animal | kg | kcal |
|---|---|---|
| Mouse | 0.021 | 3.6 |
| Rat | 0.282 | 28.1 |
| Guinea pig | 0.410 | 35.1 |
| Rabbit | 2.980 | 167.0 |
| Cat | 3.000 | 152.0 |
| Macaque | 4.200 | 207.0 |
| Dog | 6.600 | 288.0 |
| animal | kg | kcal |
|---|---|---|
| Goat | 36.0 | 800 |
| Chimpanzee | 38.0 | 1090 |
| Sheep ♂ | 46.4 | 1254 |
| Sheep ♀ | 46.8 | 1330 |
| Woman | 57.2 | 1368 |
| Cow | 300.0 | 4221 |
| Young cow | 482.0 | 7754 |
First plot: Linear
plot(kcal ~ kg, data=kleiber)
Second plot: semi-log
plot(log(kcal) ~ kg, data=kleiber)
Third plot: log-log
plot(log(kcal) ~ log(kg), data=kleiber)
Which one seems more “straight”?
The plot that seems more straight line is the log-log plot
Therefore we need a log-log model.
model <- lm(log(kcal) ~ log(kg), data=kleiber) coef(model)
(Intercept) log(kg)
4.206 0.756
What is the interpretation of these coefficients?
If \[\log(kcal)=4.21 + 0.756\cdot \log(kg)\] then \[kcal=\exp(4.21) \cdot kg^{0.756} =67.1 \cdot kg^{0.756}\]
Therefore:
- For a 1kg animal, the average energy consumption is \(\exp(4.21) = 67.1\) kcal
- The energy consumption increases at a rate of \(0.756\) kcal/kg.
This is Kleiber’s Law
“An animal’s metabolic rate scales to the ¾ power of the animal’s mass”.
Google it
This is different from previous class
Two ways to do logarithmic scales
Depending on the goal, we use different versions of semi-log and log-log plots
For understanding the data, we do
plot(log(kcal) ~ kg, data=kleiber)
For publishing in a paper, we do
plot(kcal ~ kg, data=kleiber, log="y")
Example
plot(log(kcal) ~ kg,
data=kleiber)
plot(kcal ~ kg,
data=kleiber, log="y")
Example
plot(log(kcal) ~ log(kg),
data=kleiber)
plot(kcal ~ kg, data=kleiber, log="xy")
Using the model to predict
What can we do with the model?
Models are the essence of scientific research
They provide us with two important things
- An explanation for the observed patterns of nature
- A method to predict what will happen in the future
Predicting with the model
predict(model, newdata)
where newdata is a data frame with column names corresponding to the independent variables
If we omit newdata, the prediction uses the original data as newdata
predict(model) == predict(model, newdata=data)
Results
What is wrong here?
| animal | kg | kcal | predicted |
|---|---|---|---|
| Mouse | 0.021 | 3.6 | 1.28 |
| Rat | 0.282 | 28.1 | 3.25 |
| Guinea pig | 0.410 | 35.1 | 3.53 |
| Rabbit | 2.980 | 167.0 | 5.03 |
| Cat | 3.000 | 152.0 | 5.04 |
| Macaque | 4.200 | 207.0 | 5.29 |
| Dog | 6.600 | 288.0 | 5.63 |
| animal | kg | kcal | predicted |
|---|---|---|---|
| Goat | 36.0 | 800 | 6.92 |
| Chimpanzee | 38.0 | 1090 | 6.96 |
| Sheep ♂ | 46.4 | 1254 | 7.11 |
| Sheep ♀ | 46.8 | 1330 | 7.11 |
| Woman | 57.2 | 1368 | 7.26 |
| Cow | 300.0 | 4221 | 8.52 |
| Young cow | 482.0 | 7754 | 8.88 |
Undoing the logarithm
We want to predict the metabolic rate, depending on the weight
The independent variable is \(kg\), the dependent variable is \(kcal\)
But our model uses only \(\log(kg)\) and \(\log(kcal)\)
So we have to undo the logarithm, using \(\exp()\)
Correct formula for prediction
predicted_kcal <- exp(predict(model))
| animal | kg | kcal | predicted |
|---|---|---|---|
| Mouse | 0.021 | 3.6 | 3.62 |
| Rat | 0.282 | 28.1 | 25.76 |
| Guinea pig | 0.410 | 35.1 | 34.19 |
| Rabbit | 2.980 | 167.0 | 153.11 |
| Cat | 3.000 | 152.0 | 153.89 |
| Macaque | 4.200 | 207.0 | 198.46 |
| Dog | 6.600 | 288.0 | 279.29 |
| animal | kg | kcal | predicted |
|---|---|---|---|
| Goat | 36.0 | 800 | 1007 |
| Chimpanzee | 38.0 | 1090 | 1049 |
| Sheep ♂ | 46.4 | 1254 | 1220 |
| Sheep ♀ | 46.8 | 1330 | 1228 |
| Woman | 57.2 | 1368 | 1429 |
| Cow | 300.0 | 4221 | 5001 |
| Young cow | 482.0 | 7754 | 7157 |
Visually
plot(log(kcal) ~ log(kg), data=kleiber) lines(predict(model) ~ log(kg), data=kleiber)
## Visually
plot(kcal ~ kg, data=kleiber, log="xy") lines(exp(predict(model)) ~ kg, data=kleiber)
In the paper
Moore’s Law
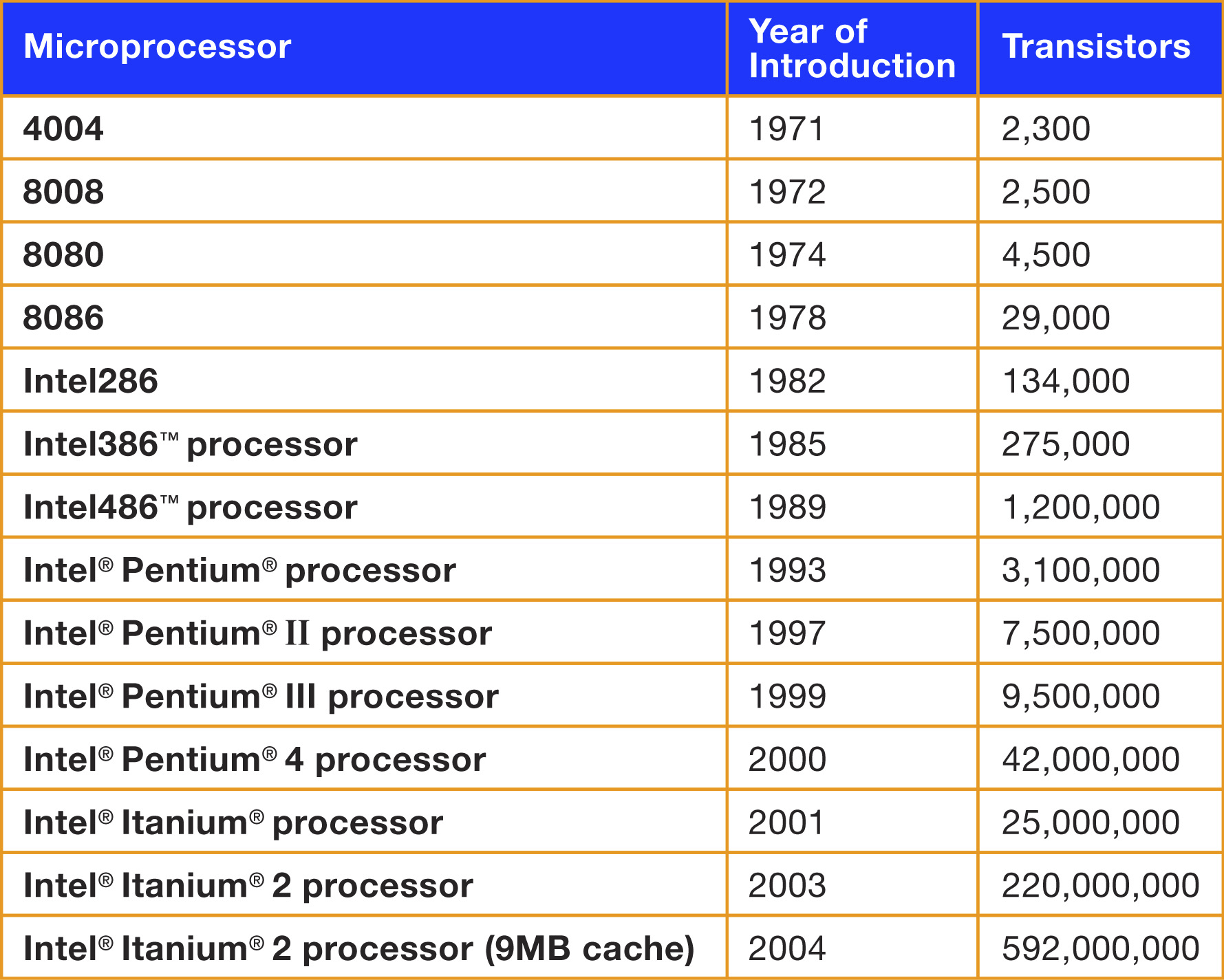
Real data: Number of transistors in chips v/s year
plot(count~Date, data=trans)
Semilog scale: Number of transistors v/s year
plot(log(count) ~ Date, data=trans)
Semi-log means exponential growth
we have straight line on the semi-log
That is, log(y) versus x \[\log(y)=log(I)+log(R)\cdot x\] In this case the original relation is \[y=I\cdot R^x\]
Model
model <- lm(log(count) ~ Date, data=trans) exp(coef(model))
(Intercept) Date 7.83e-295 1.41e+00
Semi-log scale and model
plot(count ~ Date, data=trans, log="y") lines(exp(predict(model)) ~ Date, data=trans)
Meaning
Every year processors grow by a factor of
exp(coef(model)[2])
Date 1.41