Class 1: Why do we care about Bioinformatics?
Bioinformatics
Andrés Aravena
October 5, 2023
Welcome to “Bioinformatics”
Today’s ideas
- What “Bioinformatics” is and is not
- Why you should care
- How to get bioinformatic data for free
- What kind of data we can get
- What is important in the data
Bioinformatics
what it is and what it isn’t
Molecular Biology 101
- DNA
- RNA
- Proteins
- Metabolism
What is Bioinformatics?
- Genomics
- sequences of DNA, RNA, AA
- Transcriptomics
- gene’s expression
- Proteomics
- 3D structure and interactions
- Metabolomics
- metabolites
What Bioinformatics is not?
- Using computers in a hospital
- Handling patient information
- Laboratory Information Management
- Microscope image analysis
Big picture
for this course, according to
“Bioinformatics Core Competencies for
Undergraduate Life Sciences Education.”
Sayres, et al. PLoS ONE 13, no. 6 (2018): 1–20.

Genomics
- DNA sequencing
- Pairwise Alignment
- Multiple Alignment
- Genome Assembly
- Primer design
- Finding Binding Sites
Transcriptomics
Measuring gene expression
- qPCR
- Microarrays
- RNAseq
Mostly about statistics
Proteomics
- Find protein sequence
- mass spectrometry
- Find protein structures
- X-ray diffraction analysis
- Computational Biology prediction
- Find protein-protein interactions
What we should do here
- Role
- Concepts
- Statistics
- Access
- Tools
- Pathways
- Metagenomics
- Scripting
- Software
- Computational environment
Sayres, et al. “Bioinformatics Core Competencies for
Undergraduate Life Sciences Education.”
PLoS ONE 13, no. 6 (2018): 1–20. https://doi.org/10.1371/journal.pone.0196878.
Role
Understand the role of computation and data mining in hypothesis-driven processes within the life sciences
Concepts
Understand computational concepts used in bioinformatics
- meaning of algorithm
- bioinformatics file formats
Statistics
Know statistical concepts used in bioinformatics
- E-value
- z-scores
- t test
- type-1 and type-2 error
Access genomic data
Know how to access genomic data
- NCBI nucleotide databases
- EBI
Use genomic Tools
Be able to use bioinformatics tools to analyze genomic data
- BLASTN
- genome browser
Access expression
Know how to access gene expression data
- UniGene
- GEO
- SRA
Tools expression
Be able to use bioinformatics tools to analyze gene expression data
- GeneSifter
- David
- ORF Finder
Access proteomic data
Know how to access proteomic data
- NCBI protein databases
Tools proteomic
Be able to use bioinformatics tools to examine protein structure and function
- BLASTP
- Cn3D
- PyMol
Access metabolomic
Know how to access metabolomic and systems biology data
- Human Metabolome Database
Pathways
Be able to use bioinformatics tools to examine the flow of molecules within pathways/networks
- Gene Ontology
- KEGG
Metagenomics
Be able to use bioinformatics tools to examine metagenomics data
- MEGA
- MUSCLE
Scripting
Know how to write short computer programs as part of the scientific discovery process
- write a script to analyze sequence data
Software
Be able to use software packages to manipulate and analyze bioinformatics data
- Geneious
- Vector NTI Express
- spreadsheets
Computational environment
Operate in a variety of computational environments to manipulate and analyze bioinformatics data
- Mac OS, Windows
- web- or cloud-based
- Unix/Linux command line
What we really do here
We focus on How to understand results
- Role: What is bioinformatics
- Access: using NCBI, EBI
- Concepts: file formats and more
- Tools: understanding tools output
- Statistics: E-values, error type-1 and type-2
More Concepts
- Pairwise Alignment
- Global
- Semi-global
- Local
- Multiple Alignment
- Cost
- Heuristics
- Trees
- Taxonomy
- Phylogenetic
- Ontology
Practical details
Course’s blog
My blog is at https://www.dry-lab.org/
Course’s blog at https://www.dry-lab.org/blog/2023/bioinfo/
All material will be published there
Diagnostic quiz

Why you should care
about bioinformatics
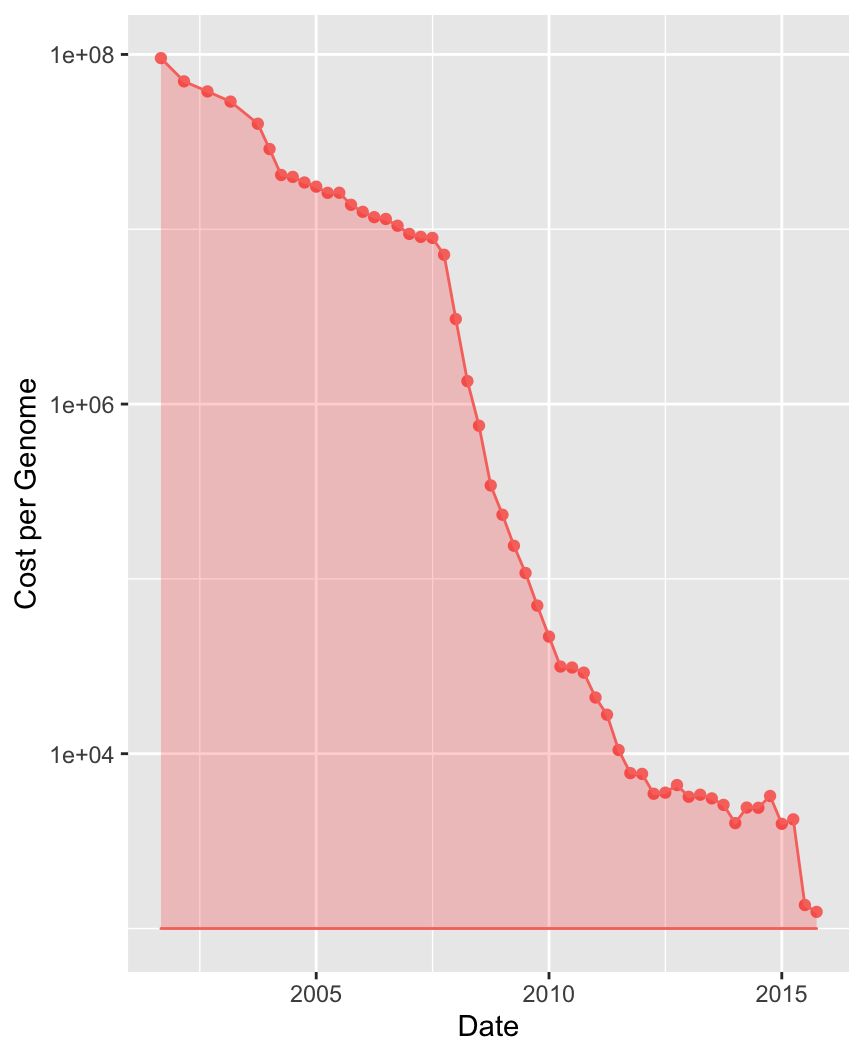
Technology changes fast
In 2001, the cost of sequencing the first human genome was USD 108
Today you can have your own genome for 1000 USD
The problem is no longer how to do the experiment
Instead is how do we make sense of the results
Manual jobs are now done by computers
Will a robot replace you?
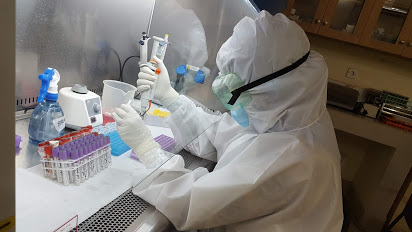

We need data
International Nucleotide Sequence Database Collaboration
There are three large data repositories
- National Center for Biotechnology Information, NCBI
- National Library of Medicine
- National Institutes of Health, USA
- National Library of Medicine
- European Bioinformatics Institute, EMBL-EBI
- European Molecular Biology Laboratory
- DNA Data Bank of Japan (DDBJ)
- National Institute of Genetics (NIG) Japan
They all have the same data
These three databases interchange all sequence data
but they may have different structure
All data is available for free
Research payed with public money must be uploaded here
Good journals also require to upload data
The NCBI website
GenBank and RefSeq
- GenBank
- genetic sequence database, an annotated collection of all publicly available DNA sequences. Anybody can upload directly.
- RefSeq
- curated subset of GenBank
DNA Databases
- Nucleotide
- most of the sequence data from GenBank, except environmental
- SRA
- sequencing data from the next generation sequencing platforms
Protein Databases
- Protein
- amino acid sequences from the translations of coding regions provided on nucleotide records in GenBank, also imported from the outside data sources (PIR, UniProtKB/Swiss-Prot, Protein Data Bank)
- Protein Clusters
- collection of related protein sequences (clusters) consisting of Reference Sequence proteins encoded by complete prokaryotic genomes, eukaryotic organelle plasmids and genomes.
Protein Databases
- Conserved Domains
- protein domains represented by sequence alignments and profiles for protein domains conserved in evolution. It includes alignments of the domains to known three-dimensional protein structures.
- Structure
- Molecular Modeling Database (MMDB) contains experimental data from crystallographic and NMR structure determinations. The data for MMDB are obtained from the Protein Data Bank (PDB)
Gene Expression Omnibus
- GEO Datasets
- gene expression data sets from the Gene Expression Omnibus (GEO) repository of microarray data
- GEO Profiles
- individual gene expression profiles assembled from GEO
- Probe
- nucleic acid reagents designed for use in a wide variety of biomedical research applications including genotyping, gene expression studies, SNP discovery, genome mapping, and gene silencing
Databases
- Taxonomy
- names and phylogenetic lineages of the more than 350,000 species that have molecular data in the NCBI databases
- MeSH (Medical Subject Headings)
- controlled vocabulary and classification system (ontology) used for indexing articles in PubMed. MeSH terminology provides a consistent way to retrieve information that may use different terminology for the same concepts
Literature
- PubMed
- database of citations and abstracts for biomedical literature from MEDLINE and additional life science journals
- PubMed Central
- (PMC) is the U.S. National Library of Medicine's digital archive of life sciences journal literature. PMC contains full-text manuscripts deposited by authors or articles provided by the publisher
Books
- Bookshelf
- full-text books that can be searched online and that are linked to PubMed records
- NCBI Web Site Search
- database of static NCBI web pages, documentation, and online tools
- NLM Catalog
- records for books, journals, audiovisuals, computer software, electronic resources, and other materials in the National Library of Medicine (NLM) collections
Databases
- Assembly
- genome assemblies. The same genome can have several versions
- Gene
- genes from completely sequenced genomes and that have an active research community to contribute gene-specific data
- Genome
- sequence and map data from the whole genomes. They represent both completely sequenced genomes and those with sequencing in-progress
Databases
- BioProject
- complete and incomplete (in-progress) large-scale molecular projects including genome sequencing and assembly, transcriptome, metagenomic, annotation, expression and mapping projects.
- BioSample
- contains descriptions of biological source materials used in studies that have data in other NCBI molecular databases such as Assembly, Nucleotide and SRA.
Other databases
- MedGen
- ClinVar
- OMIM
- PopSet
- PubChem BioAssay
- PubChem Compound
- PubChem Substance
- UniGene
- GTR
- BioSystems
Searching into NCBI
“Clipboard” and “My Collections”
The Clipboard is a temporary place on the NCBI website to save records.
- limited to 500 items on each database
- lost after eight hours of inactivity
My Collections that is a part of the My NCBI service is a more permanent place to save records.
You need to create an NCBI account to use My NCBI. It is easy and free
Pre-computed answers
There are two major kinds of relationships in the NCBI website:
- computationally derived associations within a database (neighbors)
- relationships based on information present on the records themselves (hard links)
Combining neighbors and hard links can be an especially effective method for navigating across data and finding the most useful information
Queries
NCBI Entrez queries
Searching NCBI has much more options than Google
(do you know Google options?)
By default the query text is searched in any part of any database
But you can specify the fields where you are looking for
- Title of a paper
- author
- date
- taxonomic id
Entrez Examples
protease NOT hiv1[organism]- This will limit the search to all proteases, except those in HIV 1.
1000:2000[slen]- This limits the search to entries with lengths between 1000 to 2000 bases for nucleotide entries, or 1000 to 2000 residues for protein entries.
Entrez Examples
Mus musculus[organism] AND biomol_mrna[properties]- This limits the search to mouse mRNA entries in the database. For common organisms, one can also select from the pulldown menu.
Entrez Examples
10000:100000[mlwt]- This limits the search to protein sequences with calculated molecular weight between 10 kD to 100 kD.
src specimen voucher[properties]- This limits the search to entries that are annotated with a /specimen_voucher qualifier on the source feature.
Entrez Examples
all[filter] NOT environmental sample[filter] NOT metagenomes[orgn]- This excludes sequences from metagenome studies and uncultured sequences from anonymous environmental sample studies
Creating advanced queries
Quotes " are important
The fields are written inside brackets []
Each database page includes an Advanced Search option
Combining queries
Entrez queries can be single words, short phrases, sentences, database identifiers, gene symbols, or names
AND: Finds documents that contain terms on both sides of the operator terms. The intersection of both searches.
OR: Finds documents that contain either term. The union of both searches.
NOT: Finds documents that contain the term on the left but not the term on the right of the operator. The subtraction of the right side from the left side
Example
AND must be in uppercase. It is recommended to also use
uppercase for OR and NOT
Operators are processed left-to-right
promoters OR response elements NOT human AND mammalsParenthesis can be used to control the evaluation order
g1p3 AND (response element OR promoter)
Dates and Other Ranges
Certain fields can accept ranges of values
- Publication Date, Modification Date, Accession, Molecular Weight, and Sequence Length
Low and high numbers are entered with a colon “:” between them followed by the field
110:500[Sequence Length] 2015/3/1:2016/4/30[Publication Date]
NCBI online documentation
We can get a different explanation in the public documentation made by NCBI
All documents made by NCBI are public domain