Türkçe bilmiyorum 😟
I am
- Assistant Professor at Istanbul University
- Mathematical Engineer, U. of Chile
- PhD Informatics, U Rennes 1, France
- PhD Mathematical Modeling, U. of Chile
- not a Biologist
- but an Applied Mathematician who can speak “biologist language”



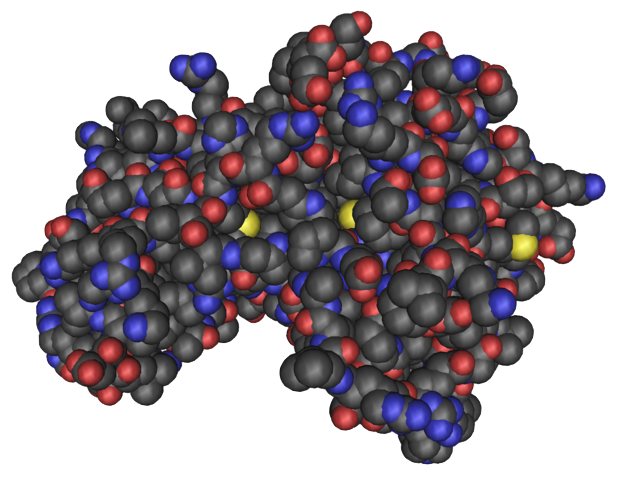
 Given the sequence of a protein
Given the sequence of a protein